DNA Sequencers
From sequencing to cure a long way to go
The ability to read genomes has transformed understanding of biology. Being able to write them would give unprecedented control over the fabric of life.
The DNA sequencers market is projected to see a steady growth in coming years. Demand for accurate, inexpensive, and fast DNA sequencing data has led to the evolution and dominance of a new generation of sequencing technologies. The use of genetic information has the potential to provide individuals and medical professionals with an arsenal of life-impacting information. In addition, different sciences are receiving benefits of this technique, including forensic sciences, molecular biology, biotechnology, genetics, anthropology, and archaeology. DNA sequencing is expected to revolutionize the conceptual foundation of these fields.
However, issues related to public health and safety could pose a challenge to the DNA sequencing market growth. The first method described for DNA sequencing was called the plus-and-minus (Sangers) method. However, owing to the inefficiency of this method, the manufacturers described a breakthrough method for sequencing oligonucleotides via enzymic polymerization. The enzymic method for DNA sequencing has been used for genomics research as the main tool to generate the fragments necessary for sequencing, regardless of the sequencing strategies. Since completion of the first human genome sequence, demand for a faster and cheaper sequencing method has increased greatly. This demand has driven the development of second-generation sequencing method or next-generation sequencing (NGS).
Currently, NGS is recognized as the primary technology responsible for the burgeoning field of oncology, Mendelian diseases, and complex disease testing. NGS assays have become an integral process in several areas of clinical diagnostics.
The all-in-one approach offered by NGS assays minimizes the time required to identify the cause of a condition, compared to sequential molecular tests, contributing to the improved health outcome. In the current scenario, liquid biopsies for early detection and monitoring of cancer and diversification of the reproductive health market exhibits substantial promise for technology in the near future.
In recent times, the miniaturization of NGS instruments with the introduction of palm-sized sequencers is one of the key developments that have driven the market. The continuous evolution of NGS technologies with respect to efficiency and cost is anticipated to boost the adoption in non-conventional applications such as agri-genomics and infectious disease testing.
The advent of liquid biopsies in cancer diagnostics is one of the most notable developments of NGS. NGS-based liquid biopsies utilize cell-free, circulating tumor DNA (ctDNA) as a noninvasive cancer biomarker for real-time cancer detection and monitoring. In addition, NGS offers high specificity and sensitivity during the detection of low-level ctDNA in the bloodstream. The targeted cancer panel sequencing or whole exome sequencing can be conducted for patient-derived cancer cells, derived both from solid tumor samples and liquid biopsies.
Recent FDA approvals of DNA home tests and the emergence of DTC genetic testing companies in developing countries like India are anticipated to drive the revenue in this segment. Mapmygenome, DNA Labs India, Positive Biosciences, Xcode, and EasyDNA are some DTC companies operating in India. These companies offer products online through their websites or e-commerce companies.
Despite opening new edge of genomics research, the fundamental shift away from the conventional sequencing to the NGS technologies has left many undiscovered applications and capabilities of these new technologies, especially those in clinical segment. In addition, the evolution of new sequencing technologies has raised the demand for commercially available platforms, whose diverse applications are restricting the potential of NGS for clinical research and physician scientists.
Indian market
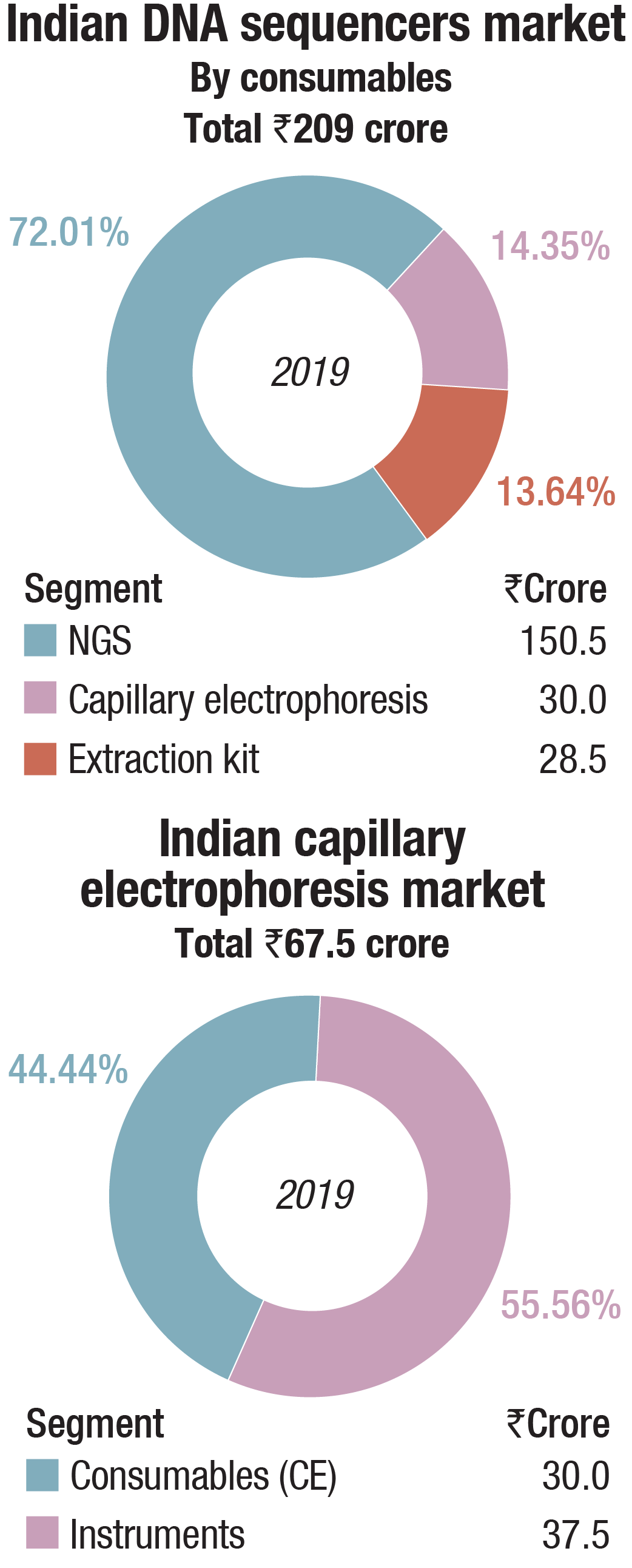
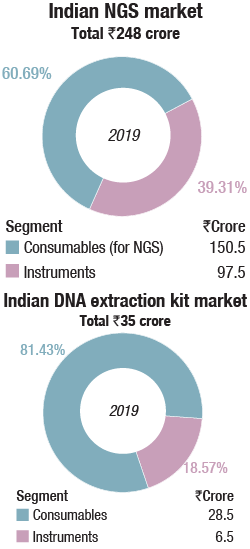
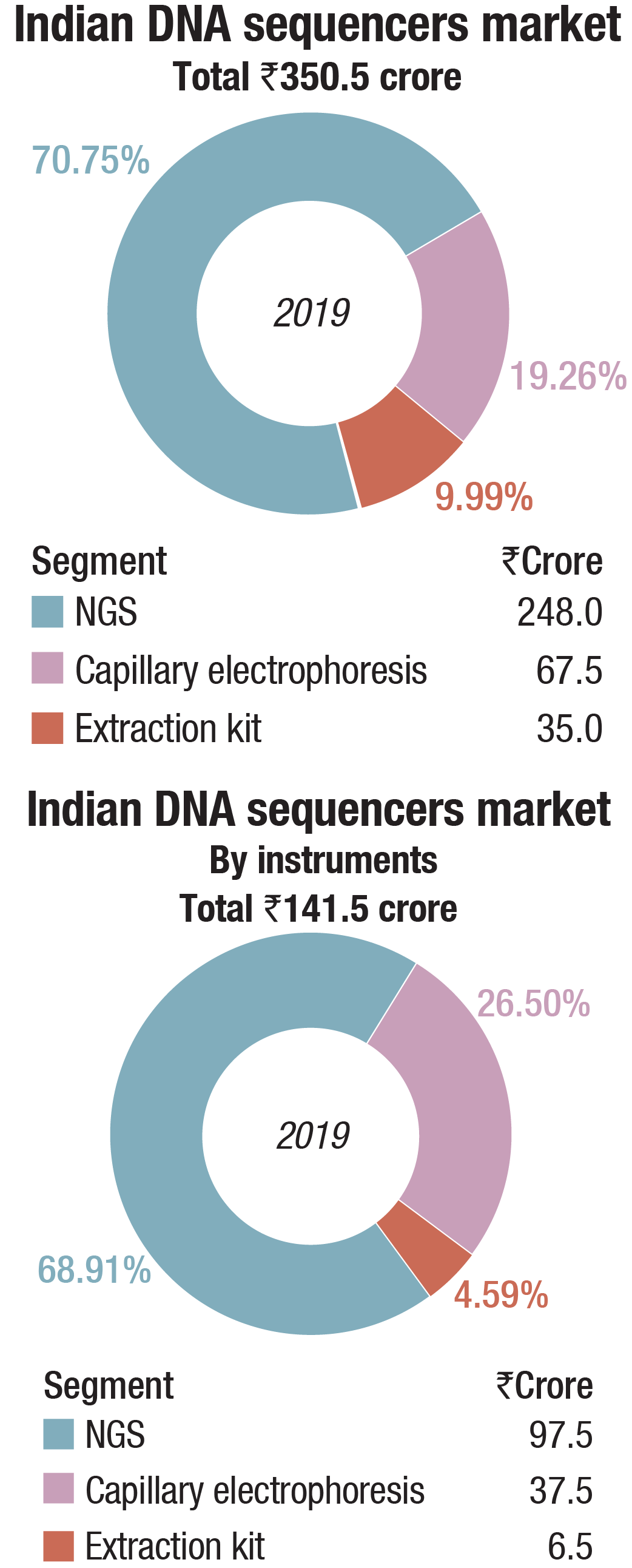 The Indian market in 2019 for DNA sequencers is estimated at Rs 350.5 crore. It may be segmented as capillary sequencers, NGS, and DNA-extraction kits. NGS continues to dominate the segment, with 69 percent share by instruments and 72 percent share by consumables, by value.
The Indian market in 2019 for DNA sequencers is estimated at Rs 350.5 crore. It may be segmented as capillary sequencers, NGS, and DNA-extraction kits. NGS continues to dominate the segment, with 69 percent share by instruments and 72 percent share by consumables, by value.
The highest adoption is in the clinical market, which in 2019 saw an increase in demand of 25 percent, primarily from the large genomic testing labs and specialized labs. The genomics space in India is estimated as a Rs 450-crore industry.
Post-COVID lockdown, genome sequence information from the virus is helping researchers identify transmission patterns of the virus in minute detail. Scientists at NIBG, GBRC, CCMG, and IGIB, apart from other observations, are keeping track to whether the A2a or the O strain gain more steam in India.
The field of genomics is getting a lot of traction from some large genome projects being undertaken by the government too.
The Genome India project, spearheaded by the Centre for Brain Research at Indian Institute of Science aims to ultimately build a grid of the Indian reference genome to understand fully the type and nature of diseases and traits that comprise the diverse Indian population. The project is a collaboration of 20 institutions, including the Indian Institute of Science and selected IITs.
 The Indian Human Microbiome Initiative Project, led by The National Centre for Microbial Resource (NCMR)–National Centre for Cell Science (NCCS)–has received funding of Rs 150 crore from the union government. The proposed initiative is a collaborative effort between 11 research institutes and universities across the country. The project will include collection of saliva, stool, and skin swabs of 20,600 Indians across 103 endogamous communities across various ethnic groups from different geographical regions. Recent technological advances in DNA sequencing and the development of meta-genomics are making it feasible to analyze the entire human microbiome.
The Indian Human Microbiome Initiative Project, led by The National Centre for Microbial Resource (NCMR)–National Centre for Cell Science (NCCS)–has received funding of Rs 150 crore from the union government. The proposed initiative is a collaborative effort between 11 research institutes and universities across the country. The project will include collection of saliva, stool, and skin swabs of 20,600 Indians across 103 endogamous communities across various ethnic groups from different geographical regions. Recent technological advances in DNA sequencing and the development of meta-genomics are making it feasible to analyze the entire human microbiome.
The Council of Scientific and Industrial Research (CSIR) has launched an ambitious project IndiGen to sequence whole genomes of diverse ethnic Indian population to develop public health technology applications. The CSIR in October 2019 announced sequencing of 1008 Indian genomes as part of the project. It aims to complete the sequencing of at least 10,000 Indian genomes over the next three years.
The area of single-cell genomics is developing at an unprecedented speed. Single-cell sequencing examines the sequence information from individual cells with optimized NGS technologies, providing a higher resolution of cellular differences and a better understanding of the function of an individual cell in the context of its microenvironment. For example, in cancer, sequencing the DNA of individual cells can give information about mutations carried by small populations of cells. In development, sequencing the RNAs expressed by individual cells can give insight into the existence and behavior of different cell types. In microbial systems, a population of the same species can appear to be genetically clonal, but single-cell sequencing of RNA or epigenetic modifications can reveal cell-to-cell variability that may help populations rapidly adapt to survive in changing environments.
 2019 also saw the setting up of a new advanced Forensic DNA Analysis Lab and a state-of-the-art DNA Analysis Centre in Central Forensic Science Laboratory, Chandigarh, under the Nirbhaya Fund scheme with an allocation of Rs 99.76 crore. This facility has an annual capacity to examine 2000 forensic cases relating to sexual assault, homicide, paternity, human identification, and mitochondrial DNA. Approval from the Ministry of Home Affairs for setting up and upgrading of DNA analysis and cyber forensic units in State Forensic Science Laboratories in 13 States/Union Territories under the Nirbhaya Fund scheme was also received. Apart from this, in the fiscal year, 25 states had been given an allocation of Rs 108.40 crore for strengthening the forensic science facilities, including DNA analysis facilities, as part of annual State Action plan under the scheme for modernization of police forces. The Directorate of Forensic Science Services has six central forensic science laboratories (CFSLs), situated at Bhopal (Madhya Pradesh), Chandigarh, Guwahati (Assam), Hyderabad (Telangana), Pune (Maharashtra) and Kolkata (West Bengal).
2019 also saw the setting up of a new advanced Forensic DNA Analysis Lab and a state-of-the-art DNA Analysis Centre in Central Forensic Science Laboratory, Chandigarh, under the Nirbhaya Fund scheme with an allocation of Rs 99.76 crore. This facility has an annual capacity to examine 2000 forensic cases relating to sexual assault, homicide, paternity, human identification, and mitochondrial DNA. Approval from the Ministry of Home Affairs for setting up and upgrading of DNA analysis and cyber forensic units in State Forensic Science Laboratories in 13 States/Union Territories under the Nirbhaya Fund scheme was also received. Apart from this, in the fiscal year, 25 states had been given an allocation of Rs 108.40 crore for strengthening the forensic science facilities, including DNA analysis facilities, as part of annual State Action plan under the scheme for modernization of police forces. The Directorate of Forensic Science Services has six central forensic science laboratories (CFSLs), situated at Bhopal (Madhya Pradesh), Chandigarh, Guwahati (Assam), Hyderabad (Telangana), Pune (Maharashtra) and Kolkata (West Bengal).
Some large NGS customers include National Institute of Biomedical Genomics (NIBMG), All India Institute of Medical Sciences (AIIMS), Indian Institute of Science, Bangalore, The National Institute of Plant Genome Research, and Tata Memorial Hospital and MedGenome Labs Ltd. The customers for capillary sequencers also include the lab chains as Metropolis and Lal Pathlabs Labs.
Global market
The global DNA sequencing market is projected to grow to USD 17.6 billion by 2025, driven by a compounded annual growth rate of 18.1 percent. With the reducing cost of sequencing, diagnostics segment displays the potential to grow at over 18.5 percent, estimates ResearchAndMarkets. The shifting dynamics supporting this growth makes it critical for businesses in this space to keep abreast of the changing pulse of the market. Poised to reach over USD 6.7 billion by the year 2025, diagnostics will bring in healthy gains, adding significant momentum to global growth.
Representing the developed world, the United States will maintain a 19.5-percent growth momentum. Within Europe, which continues to remain an important element in the world economy, Germany will add over USD 667.5 million to the region’s size and clout in the next 5 to 6 years. Over USD 840.9 million worth of projected demand in the region will come from the Rest of Europe markets.
In Japan, diagnostics will reach a market size of USD 632.7 million by 2025. As the world’s second-largest economy and the new game changer in global markets, China exhibits the potential to grow at 17.7 percent over the next couple of years, and add approximately USD 3.1 billion in terms of addressable opportunity for the picking by aspiring businesses and their astute leaders.
The global NGS market size was estimated at USD 9.78 billion in 2019, and is projected to grow at a CAGR 11.7 percent during 2020–2027, predicts Grand View Research. With the reducing cost of sequencing, NGS is anticipated to witness diversification into a substantial number of clinical areas. Library preparation, assay designing, sequencing, and data analysis are the key phases of the entire NGS procedure. Out of all, sequencing is recognized as the critical phase as it demands skilled expertise to handle NGS technology. Moreover, sequencing is relatively capital-intensive. These factors have contributed to the largest share of this segment. On the other hand, there is constant growth in the development of data generated through NGS technology.
This emphasizes the need for development in data analysis algorithms and software. Owing to this factor, the data analysis segment is anticipated to be the source of lucrative growth in the coming years. The key players are collaborating and acquiring technologies as well as other companies to accelerate their data analysis workflows. Illumina has accelerated its data analysis workflows through the acquisition of Edico Genome and its Dragen Bio-IT Platform.
Academic research entities generated the highest revenue in 2019 owing to the high use of NGS platforms within universities and institutes to drive basic research programs. University of Pittsburgh, University of Minnesota, University of Chicago, and Johns Hopkins University are some key universities that employ NGS technology or offer associated services to researchers.
Entities within academic settings have collaborated with key market players, such as Illumina and Pacific Bioscience to offer NGS services for researchers.
Clinical laboratories have begun adopting NGS as a gold standard for the testing of inherited conditions owing to its analytic accuracy, high throughput, and cost-effectiveness. NGS-based clinical genetic tests have been implemented for CNV analysis and variant detection on a broad scale at clinical laboratories. This is expected to positively influence the future revenue growth of the NGS market for hospitals and clinics.
Thermo Fisher Scientific Inc., F. Hoffmann-La Roche Ltd., and Illumina, Inc. are projected to lead in the forthcoming period owing to their rising number of strategic partnerships, robust geographical presence, and diversified product portfolio. Other companies are also striving persistently to gain the maximum NGS market share by launching new products. With the growing clinical application of NGS, the companies are collaborating with clinical research entities to decipher the pathogenesis of various diseases, using NGS platforms. Moreover, the companies are undertaking business strategies to drive down the NGS cost to nearly USD 100.
Some of the important key industry developments include:
June 2019. Saphetor SA, a precision medicine company headquartered in Switzerland, collaborated with Swift Biosciences Inc., a developer of molecular biology reagents for research and diagnostic applications, based in Michigan, to provide a set of NGS technologies and products for clinical, translational, and genomics research applications.
March 2019. Illumina collaborated with the Lundbeck Foundation GeoGenetics Centre at the University of Copenhagen, Denmark, to establish the link between infectious pathogens and neurological and mental disorders.
January 2019. PierianDx, Inc., a provider of a comprehensive combination of technology and services, based in Missouri, declared its multi-year, non-exclusive, and strategic partnership with Illumina, Inc., a developer of integrated systems, headquartered in California. The main aim of this collaboration is to offer a variant interpretation and reporting solution based on the former’s clinical genomics knowledgebase and clinical genomics workspace platform for specific Illumina oncology products.
Technological advances in genome sequencing
The Human Genome Project, which aimed to sequence the human genome in its entirety, was declared complete in 2003, and was celebrated as a major milestone for science. Genetic sequencing tools had gotten more sophisticated, efficient, and inexpensive, which helped make the achievemen t possible. But behind the scenes, there were some problems. The human genome contains many repetitive regions that do not code for genes and can be extremely challenging to sequence, so there were some gaps. Most of the work was also performed using a genomic sample from one individual, so it was an incomplete picture. The 1000 Genomes Project, completed in 2012, sequenced 1092 genomes so that we would learn more about variation in the human genome. But that is still an underrepresentation and the data may not be included in reference sequences, potentially because of quality issues.
Last year, the National Institute of Health announced an initiative to address these problems. Called the Human Pangenome Reference, the project aims to sequence the entire genomes of 350 individuals.
“One human genome cannot represent all of humanity. The human pangenome reference will be a key step forward for biomedical research and personalized medicine. Not only will we have 350 genomes representing human diversity, they will be vastly higher quality than previous genome sequences,” said David Haussler, professor of biomolecular engineering at UC Santa Cruz and director of the UC Santa Cruz Genomics Institute.
Nobel Laureate Frederick Sanger and colleagues created Sanger sequencing for reading a genetic sequence in 1977. After a DNA sequence has been amplified, each base is labeled and then read from one end. In 1995, pairwise end sequencing was shown to be useful for sequencing whole genomes, and the technology was used to sequence the human genome.
A library of fragments is prepared from a genome, and the fragments are read. Computational tools assemble the fragments into longer sequences. Nanopore sequencing, which pushes a molecule through a really tiny pore, detecting the base as it moves through, was created as one of the third-generation sequencing technologies.
With an intensive effort, including 150,000 hours of commuting time, researchers at UC Santa Cruz were able to use nanopore technology for long-read human genome sequencing. Around a year later, the cost was brought down significantly, and the results were then obtained within about a week. “We sequenced eleven human genomes in nine days, which was unprecedented at the time,” said UC Santa Cruz Research Scientist Miten Jain.
It has now been taken even further; an algorithm has been designed that can use long-read sequencing data to assemble as complete human genome in around six hours, and for about USD 70.
The researchers said they hope their assembler will increase the pace of genomics research, and open opportunities. This includes enabling pangenome research to represent the true scale of human diversity, a decidedly more practical pursuit.
“Our new assembler was designed to be cheap and quick, with the goal to be on the cloud,” said UC Santa Cruz’s Benedict Paten. “It gives us the power to scale nanopore sequencing. Now, I am confident that we’ll be easily assembling hundreds of de novo genomes in the next couple of years.”
Writing whole genomes from scratch
The ability to read genomes has transformed understanding of biology. Being able to write them would give unprecedented control over the fabric of life.
Rapid advances in DNA sequencing and gene-editing technology mean the industry is now truly in the age of genomics. For a few hundred dollars, genetic testing companies will give a detailed rundown of ancestry, and susceptibility to a host of diseases. The first genetically modified humans are about to turn one.
The advent of CRISPR in particular has given the ability to tweak DNA with incredible precision, but still largely restricted to switching specific genes on and off, or swapping one gene for another. The field of synthetic biology wants to change that by bringing engineering principles to biology.
But there is a long way to go, and a group of leading geneticists have now laid out the technology roadmap required to get there. These are the four areas where one needs to step up their game.
Genome design. The ultimate goal of genetic modification is to produce a change in the phenotype–the outward characteristics–in the target organism. But most complex traits are the result of a complicated interplay between multiple genes and an organism’s environment, so mapping how DNA tweaks will translate into desired attributes is challenging.
Large-scale genome design will require computer programs that can do this accurately and efficiently. While projects like Synthetic Yeast 2.0 have made the first steps in this direction, the field needs to build complex new models that can predict the results of changes to the genome sequence.
Major players* in Indian DNA sequencers market–2019 |
|||
|---|---|---|---|
| Segment | Instruments | Consumables | |
| Capillary Electrophoresis | Tire I | Thermo Fisher | |
| Tire II | Thermo Fisher (brand Applied Biosystems) | NimaGen, Qiagen, GE, Millipore, Chinese brands, and home brewed labs | |
| NGS | Tier I | Illumina (Premas Life Sciences) | |
| Tier II | Thermo Fisher, Pacific Biosciences (Spinco), and Agilent | ||
| DNA Extraction Kit | Tier I | Qiagen | |
| Tier II | Thermo Fisher and Promega | Tulip and Transasia | |
| *Vendors are placed in different tiers on the basis of their sales contribution to the overall revenues of the Indian DNA sequencers market. | |||
| ADI Media Research | |||
These could still be decades away, but using machine learning to mine the wealth of biology data in public databases could accelerate them. Programs that can automate the design of experiments to cut down the number of rounds of design will also be required, as will the adoption of common data standards to enable collaboration.
DNA synthesis. Experts have been able to synthesize DNA for decades, but the most common approach is restricted to short sections of DNA, just a few hundred base pairs long. Building entire genomes requires long sequences of several thousand base pairs; so currently scientists rely on a laborious and error-prone process of stitching many smaller DNA sections together.
Large-scale genome engineering will require much faster, cheaper, and more efficient methods for DNA assembly. One nearer-term possibility is designing new enzymes that can reduce the number of errors and, therefore, boost the yield of the process. But in the long run, new technologies that can produce long and accurate sequences offer far more potential, and there are some promising enzyme-based approaches that may fit the bill.
Genome editing. While gene-editing process has come a long way, experts still struggle to make widespread changes to a genome simultaneously. If one could develop this capability, it could significantly decrease the amount of time it takes to modify organisms and even sidestep the need to build genomes from scratch.
This will mean finding ways to prevent the multitude of guide RNAs (the homing devices that tell CRISPR where in the genome to make changes) required for simultaneous edits to multiple genes from interfering with each other.
It will also be necessary to create libraries of tools for making changes across the genome, and accessibility maps that highlight how efficiently different targets can be altered. These will make it easier for scientists to plan where to make changes to achieve their desired results, and form the basis of predictive computer models that can streamline the process.
Chromosome construction. DNA is more than just a string of genes; it is packaged into chromosomes, whose number and shape vary across species. The ability to assemble and manipulate these chromosomes is still rudimentary.
Most efforts so far have relied on yeast and it has been able to deal with viral, bacterial, yeast, and algal chromosomes, as well as fragments of mice and human genomes. But engineering more specialized artificial chromosomes looks to be beyond yeast; so, there is a need to find newer, more flexible organisms that can do this.
Transplanting these chromosomes into the target organism is also a major bottleneck. Techniques like cell fusion and microinjection show promise but require funding for multidisciplinary work to bridge the gap between microfluidics research and molecular biology. There is also a need for greater understanding of the fundamental forces that govern the architecture of chromosomes and how they interact.
Outlook
A number of promising signs are pointing to a new, patient-empowered future. However, achieving all this could take decades, and will require the same kind of massive cross-disciplinary effort seen in the Human Genome Project. It will also require concerted government funding and close involvement of the private sector, if it is going to become a reality.
Second Opinion
An accurate analytical tool
Prof Nagendra Singh
National Professor, Dr BP Pal Chair,
NRCPB, ICAR New Delhi
DNA sequencing has emerged as a highly accurate analytical tool for the discovery and analysis of structural variations among different individuals of a population or species. DNA sequencing is useful for disease diagnosis, predisposition to genetic disorders, parental disputes and criminal cases, identity, and purity testing of products, based on quantitative analysis of sequence tags. The cost of sequencing has come down drastically owing to rapid developments in sequencing technology, starting from the use of radioactively labelled nucleotides in the chain-termination techniques, described by Sanger in 1977. An automated capillary electrophoresis-based sequencer from ABI and Pharmacia, using fluorescently labelled dideoxy nucleotides for the detection of sequence ladder, was the first major breakthrough in reducing the time and cost of sequencing during 1990s, which powered the early model genome sequencing projects.
However, it still required purified cloned DNA fragments as sequencing templates; hence, the cost per data point was still quite high. The advent of NGS technologies during 2005–2008, where the cloning in bacterial host cells was replaced by PCR amplification of DNA fragments on a solid support allowing massive parallel sequencing of thousands of DNA fragments, thus drastically reduced the cost per data point.
There were three independent sequencing platforms, Illumina Solexa, 454 pyrosequencing, and ABI SOLid. The last two of these have become obsolete now, but the Illumina sequencers, using fluorescently labelled reversible terminators, are still widely used and provide very high throughput and lowest cost per data point. Sequencing of pooled bar-coded samples further reduces the cost per sample and per data point.
For the last five years, we have the third-generation sequencers where even PCR amplification of the DNA template is not necessary. These sequencers are based on single-molecule real-time sequencing of DNA fragments directly isolated from the cell. These are represented by Pacific Biosciences (PacBio) and Oxford Nanopore sequences, which generate sequence reads of several Kbp (average >10kbp). The PacBio system is based on real-time imaging of fluorescently labelled deoxy nucleotides as they get incorporated in the growing polynucleotide chain during synthesis, using a single molecule of template DNA and single-molecule DNA polymerase in nanosized well (ZMW). The Oxford Nanopore system is based on measurement of change in voltage while passing of a group of five nucleotides of single-stranded DNA through a nanosize made of protein. Potentially, the Nanopore system is quite simple, using basic instrument of the size of a cellphone that plugs into your laptop and needs least number of reagents, but the sequence errors are still quite high due to pore-to-pore variations. Therefore, at present PacBio Sequel II has an edge over the Nanopore system, but the accuracy of the latter is improving rapidly, which is going to make the future sequencing very fast and cheap.
 Debjani Saha
Debjani Saha
Group Product Manager,
Premas Lifesciences
The NGS market has grown exponentially over the last decade or so, and is poised to see the biggest adoption in clinical diagnostics and precision medicine. The increased knowledge of the underlying biology of different diseases and the pathways associated has been unearthed by NGS, and has also led to tailoring of targeted therapies, which again need to be accompanied by NGS-based diagnostics. While adoption has been slow in India, due to non-reimbursement policies, enhanced clinician awareness and a collaboration between academic and private institutions will help in developing cost-effective diagnostic tools for patients.