Mass Spectrometers
MS Advances To Conquer Analytical Challenges
Future innovations in technology and instrumentation will drive novel clinical applications of MS to the forefront.
The use of mass spectrometry (MS) is becoming increasingly important for biomedical research and for clinical applications. In the last 20 years, MS has evolved from purely academic instrumentation to a technique now present in most analytical laboratories. Together with ongoing improvements in seamless software workflows and capabilities, increases in sensitivity and resolution are key drivers for this development. With the advent of the genetic revolution, the proteomic revolution has followed in close succession. The constant development of MS analytical science has pushed boundaries in the past 20 years. The high throughput required by the bio/pharma industries means that one needs not only ever increasingly precise and sensitive systems for quantitative and qualitative analyses, but also speed in acquiring the data. The data also needs to be as comprehensive as possible, providing meaningful information that can be linked with physiology and disease mechanisms. This is imperative, not only for drug discovery in bio/pharma, but also in basic life sciences research, where investigating disease biomarkers and treatment targets is fundamental. The sensitivity of mass spectrometers and increasingly more sophisticated bioinformatics tools are opening up this field to untold possibilities for biomedical researchers and clinicians. In addition, it is these novel discoveries and ongoing R&D that is facilitating the realization of precision and personalized medicine today, and in the near future.
Indian market
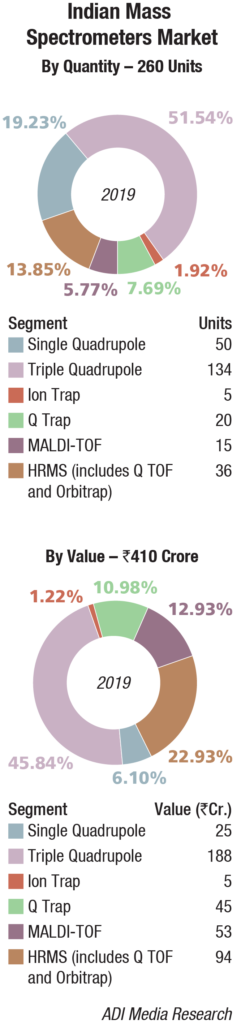 The Indian mass spectrometers market is estimated at Rs 410 crore in 2019 with 260 units. 2019 saw major price erosion in this segment with poor credit facilities available and, thus, low allocation of funds for capital equipment. If this situation continues and pharma companies stay away from investing, downtime will be huge if this vacuum-based system goes down, as much advanced preparation is required and a proper solvent that does not contaminate the system is mandatory.
The Indian mass spectrometers market is estimated at Rs 410 crore in 2019 with 260 units. 2019 saw major price erosion in this segment with poor credit facilities available and, thus, low allocation of funds for capital equipment. If this situation continues and pharma companies stay away from investing, downtime will be huge if this vacuum-based system goes down, as much advanced preparation is required and a proper solvent that does not contaminate the system is mandatory.
n of analytes to the nearest 0.001 atomic mass units. Examples of HR-MS instruments are time-of-flight (TOF) and Fourier transform ion cyclotron resonance (FTICR), which forms the basis of the orbitrap technology. Generally, these instruments measure the exact mass of analytes without fragmentation, however, they can be combined with a quadrupole in which case fragmentation is also possible and can add more selectivity to the method. However in India, given the nature of use, with focus on generic chemicals and barely any innovation done; and price points being of utmost importance, triple quad continues to be popular. The clinicians are looking to only perform targeted quantitation with ultimate sensitivity being important. Hence, HR-MS may not be appreciated in India, procured largely to meet regulatory compliance.
While MALDI-TOF is being considered archaic by some countries, as a result of its being inexpensive, easy to perform, fast and accurate, matrix-assisted laser desorption ionization, in India it continues to have relevance and has become the standard means of bacterial identification from cultures in clinical microbiology laboratories.
Government initiatives for pollution control and environmental testing are to some extent supporting the growth of this market. However, the dearth of skilled professionals and the high cost of instruments may restrain the market to a certain extent.
In 2020 a 6-7 percent growth is expected. With the coronavirus outbreak some shortage of raw material is anticipated as the formulation industry has an estimated 3-month inventory.
Globally the trend is moving toward HR-MS moving toward high res, which allows detectio
HRMS (includes Q TOF and Orbitrap)Sciex, Waters, Shimadzu (not sold), Agilent, Thermo Fisher, Brukert
Major Vendors* in Indian Mass Spectrometers Market – 2019 |
|||
| Tier I | Tier II | Tier III | Others |
| Sciex | Shimadzu, Waters, and Agilent | Thermo Fisher | Bruker, PerkinElmer, Advion, JEOL, Microbio, and Trivitron |
| *Vendors are placed in different tiers on the basis of their sales contribution to overall revenues of the Indian mass spectrometers market. | |||
| ADI Media Research | |||
Indian Mass Spectrometers Vendors Market – 2019 |
|
|---|---|
| Segment | Major Players |
| Single Quadrupole | Shimadzu, Waters, Advion, Thermo, Fisher and Agilent |
| Triple Quadrupole | Sciex, Waters, Shimadzu, and Agilent |
| Ion Trap | Thermo Fisher and Bruker |
| Q Trap | Sciex |
| MALDI -TOF & Maldi Top TOF | Bruker, Shimadzu, and JEOL |
| ADI Media Research | |
Technology trends
In the last five years or so, new MS products have addressed the full scope of user requirements through an elevated level of functionality by offering a wide range of applications as well.
Advances in MS-based strategies for probing ligand-target interactions. For the last decades, MS has been recognized as a powerful tool to study the non-covalent interactions of the ligand-target complexes with characteristics of high sensitivity, high-resolution, and high-throughput. In recent years, a number of reviews have been published on the applications of the electrospray mass spectrometry (ESI-MS) and matrix-assisted laser desorption ionization mass spectrometry (MALDI-MS) in the studies of the non-covalent complexes. It can be seen that the ESI-MS dominates the studies of non-covalent complexes. Although the high sensitivity and relative molecular mass characterization range of the MALDI-MS are generally superior to the ESI-MS for the determination of biological macromolecules, its application in the detection of non-covalent complexes is limited by sample preparation conditions and high action energy. For example, the usage of the strong acidic matrix or organic co-solvents in the ionization process is generally detrimental to the formation of the non-covalent complexes. Moreover, the presence of laser-induced polymerization and matrix adduct formation often interferes with the detection of complexes, which makes MALDI-MS relatively less useful for the studies of non-covalent complexes.
Non-covalent interactions can be classified as specificity and non-specificity, and the specific non-covalent complexes with biological functions are the focus of attention. The so-called complex peaks on the mass spectrum do not necessarily represent the actual complexes, and sometimes false positives occur. In order for the mass spectrometry data to truly reflect the non-covalent interactions of the system under test, it is necessary to use some experimental methods, such as the usage of the classical biochemical techniques of the enzymatic hydrolysis, chemical modification, etc., to change the sequences or properties of biopolymers to weaken or destroy the non-covalent bonds, and then to compare the changes in the corresponding signals on the mass spectrum before and after the reaction to infer and verify the structures of the complexes. Secondly, different sample-preparation techniques or different buffer systems need to be selected to optimize the test conditions in the experiment. It is also necessary to control the appropriate instrument conditions, especially the parameters of the ion-source part, so as to reduce the dissociation of non-covalent bonds and obtain the structural information of specific non-covalent complexes.
Proteins/enzymes play an important role in the life process, which are closely related to many malignant diseases such as tumors. Therefore, it is urgent to design and develop new small drug molecules, targeting those proteins/enzymes. The soft ionization MS-based methods could comprehensively, systematically, and accurately study the protein/enzyme-small molecule interactions, and provide information, such as the action sites (or regions) and the action intensity, so as to provide better help for drug design and disease treatment. With the development of soft ionization technologies, MS-based methods will continue to develop and improve, which will definitely promote the analysis and research of protein/enzyme-ligand interactions, so as to better serve human health.
Upcoming revolution in disease detection. Advances in mass spectrometry-based diagnostics could ignite a revolution in the field of molecular medicine. This platform has the potential to become the practical clinical analyzer of the future for nucleic acids and proteins. Mass spectrometry-based diagnostics are an example of a disruptive or nonlinear technology. Such disruptive technologies are by their very nature polarizing, causing a dynamic dichotomy of excitement as well as anxiety in the clinical diagnostic community, because this technology can potentially outperform traditional measurement and detection systems. Although major advances have been made in elucidating the genetic underpinnings of cancer, especially colorectal cancer, diagnostic methodologies for routine clinical detection and monitoring of important cancer-genetic derangements have lagged behind. Microsatellite instability (MSI), caused by mismatch repair gene silencing, is predicted to be an important early event in cancer progression. Mass spectrometry-based detection of surrogates for disease detection is a timely topic. Mass spectrometry has recently generated excitement (and anxiety) as a platform for protein-based biomarker profiling. The most reliable, sensitive, and widely available tests for the detection of cancer today are protein-ligand assays, such as ELISA systems. Finding the single disease-related protein is like searching for a needle in a haystack, requiring the separation and identification of these entities one by one.
Within the past year, a new type of protein-based diagnostic paradigm has been described – proteomic pattern diagnostics. MS-based proteomic pattern diagnostics have been used for ovarian cancer detection, and the value of this paradigm has been confirmed in other diseases, including breast and prostate cancer. What this means is that the unique tumor-host microenvironment can set off amplification cascades that may be specific to the disease process, and yet the signatures for the presence of cancer, even at its earliest stages, may be composed of untold numbers of combinations of slight but significant shifts in protein–protein interactions, protein folding, and protein abundances that are reflected in MS-based protein profiles. MS platforms, already capable of reporting tens of thousands of events in less than a few minutes from a microliter of blood, are advancing rapidly in speed, throughput, sensitivity, and on-the-fly protein identification. Semi-quantitative MSI profiling represents an additional new and exciting component of the repertoire of mass spectrometry’s diagnostic potential.
The coupling of advances in MS with adaptive and vigilant bioinformatic pattern recognition tools may dramatically change how disease is detected and monitored. The result will be a rich source of information to aid the clinician in patient management. On the basis of these nonlinear technologic advances, the clinical diagnostic landscape is now shifting dramatically. Like it or not, one may be moving beyond existing immunoassay-based (for proteins) and electrophoretic (nucleic acid) approaches. Rapid, low-cost mass spectrometry-based alternatives, with much higher clinical sensitivity and specificity for the detection and monitoring of disease, may eventually dominate clinical diagnostics. The utility and validity of this vision will be answered over the next several months to few years.
MS for metabolites and inborn errors of metabolism. Targeted MS identification of specific small-molecule metabolites in biofluids enables diagnosis of some IEMs in neonates. For instance, defects in fatty acid oxidation can be identified through acylcarnitine profiling. Other IEMs that MS can help diagnose include cystinosis, glutaric aciduria Type-I, and lysosomal disorders, which themselves span a range of illnesses, such as Fabry disease, Gaucher disease, Krabbe disease, Mucopolysaccharidoses Type-I and Type-II, Niemann-Pick-type A/B disease and C disease, and Pompe disease. A recent trend toward untargeted, systems biology utilizes metabolomics to detect alterations across the entire metabolome, which may be tied back to a mutation in a certain metabolic enzyme and hence to an IEM. This approach is not yet in clinical use but is an active area of research.
MS for cancer diagnostics and surgery. Matrix-assisted laser desorption/ionization imaging mass spectrometry (MALDI IMS) is a combination of MS with spatial information. The MS collects a spectrum across an entire histological sample, generating a spatial MS map. To date, it has been principally used to analyze cancer tissue, either for diagnostic purposes, i.e., differentiating tumor from healthy tissue, or for prognostic purposes, i.e., predicting patient survival. The method is not FDA-cleared but could be amenable to clinical use.
Way forward
The use of mass spectrometry in all areas of the pharmaceutical industry has increased markedly over the last ten years as instruments become smaller and cheaper, or smaller with increased resolution. The changes in the project portfolios across the pharmaceutical industry with novel (larger) molecules and complex drug-delivery devices means that there are many challenges where mass spectrometry will be the analytical technology of choice. However, there is also a requirement to shift to differing separation techniques in front of the mass spectrometer or for ion-mobility mass spectrometry, after the ionization has occurred. It is clear that mass spectrometry coupled to a wide range of separation technologies continues to play an essential role throughout the pharmaceutical industry, from discovery to development, to supporting a long-term supply of essential medicines to patients. The continuing evolution of MS technologies will only further strengthen the future impact and importance of MS in the healthcare and pharmaceutical industry. LC–MS is still a predominant technique and its impact will not only continue, but will be enhanced over the coming years.
Second Opinion
Recent trends and advances in mass spectrometry
 Dr Shaik Mohammad Naushad
Dr Shaik Mohammad Naushad
Head, Biochemical Genetics and Pharmacogenomics,
Sandor Speciality Diagnostics Pvt. Ltd., Hyderabad
Mass spectrometry has huge impact on all the –OMICS and has revolutionized basic and applied research in the recent years. Here, some of the most recent advances are highlighted. Vimer et al. developed a protocol using native mass spectrometry that allows rapid characterization of intact overexpressed proteins immediately from the crude samples. Using Orbitrap and quadruple time-of-flight (QTOF)-based mass spectroscopy, this protocol enables the detection of intact protein within a few hours in both prokaryotic and eukaryotic expression systems. Rapid online buffer exchange (OBE) native mass spectrometry is another new milestone achieved, which allows direct screening of structural features of pre-purified proteins, protein complexes, and clarified cell lysates. The crude sample is directly injected on to size-exclusion chromatographic column using aqueous, non-denaturing mobile phase, and then subjected electrospray ionization and mass spectrometric detection. Sample purity, the extent of oligomerization, and information of protein complex subunits can be deduced using this technology. Lohofer et al. applied matrix-assisted laser-desorption ionization (MALDI) imaging and laser ablation – inductively coupled plasma – mass spectrometry (LA-ICP-MS) imaging to visualize the localization, and measured the concentration of the MR imaging probe Gadofluorine P in the atherosclerotic plaque tissue ex vivo with a high spatial resolution. This is a commendable achievement with specific impact on the modern medicine, specifically in diagnosing coronary artery disease. Temperature-jump mass spectrometry was developed recently with capability to study kinetics (0.16–32 s) at a wide range of temperatures (10–90°C), which has enabled the study of kinetics of folding and unfolding of DNA triplex. This can be applied to study the folding and binding properties of any other biomolecule as well. Parallel accumulation-serial fragmentation (PASEF) coupled with nanoflow liquid chromatography and trapped ion mobility spectrometry (TIMS) allowed high-sensitivity lipidomics in 1 µL of human plasma. These advances facilitate deeper understanding of biological systems and better characterization of biomolecules.